
A newly identified cellular system monitors subtle variations in genetic coding, hinting at a hidden level of control over how genes are expressed.
Human DNA stores instructions for life in sequences built from just four nucleotides. Those instructions are read in three-letter groups called codons, and each codon tells the cell which amino acid to add when building a protein. Since several codons can specify the same amino acid, these alternative spellings were once thought to be mostly interchangeable.
That view has changed. Scientists now know that some of these synonymous codons help messenger RNA, or mRNA, remain stable and get translated efficiently, while others do not. In effect, two genetic sequences can encode the same protein but still behave differently inside a cell. mRNAs rich in non-optimal codons are translated less efficiently and are often broken down more quickly, but the mechanism human cells use to detect this difference has been unclear.
Researchers at Kyoto University and RIKEN, led by Osamu Takeuchi and Takuhiro Ito, set out to solve that puzzle. Their work points to a protein called DHX29 as a key part of the system cells use to read this hidden layer of genetic information.
Visualizing the Molecular Mechanism
The team began with a genome-wide CRISPR screening to search for factors involved in codon-dependent gene expression. That search identified the RNA-binding protein DHX29 as a central regulator. Follow-up RNA sequencing showed that when DHX29 is lost, mRNAs enriched in non-optimal codons accumulate, indicating that the protein normally helps keep these transcripts in check.
Using cryo-electron microscopy, the team observed how DHX29 directly interacts with the 80S ribosome. They also applied selective ribosome profiling to determine which codons were being read by ribosomes associated with DHX29.
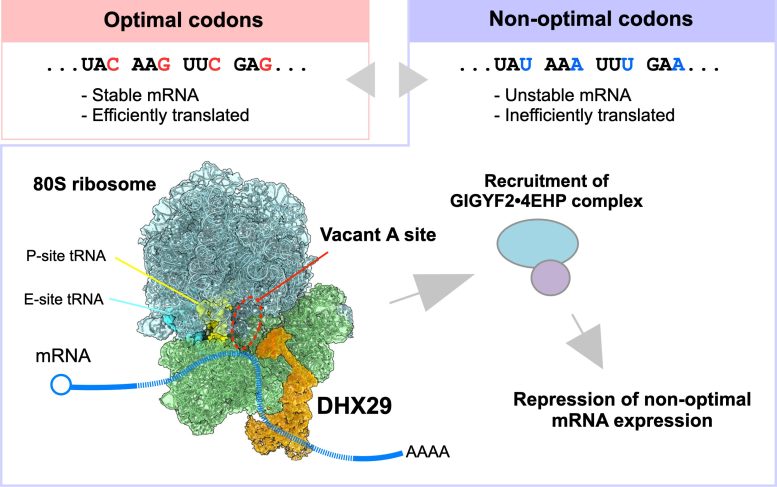
The results showed that DHX29 tends to bind ribosomes that are decoding non-optimal codons. Proteomic analyses further revealed that DHX29 recruits the GIGYF2•4EHP protein complex, which selectively suppresses mRNAs enriched in non-optimal codons.
“Together, these findings reveal a direct molecular link between synonymous codon choice and the control of gene expression in human cells,” says co-corresponding author Masanori Yoshinaga.
Broader Implications for Biology and Disease
These findings change how scientists understand the role of codon choice in regulating gene expression in human cells. The DHX29-driven mechanism may influence important biological processes such as cell differentiation, cellular homeostasis, and cancer development, suggesting wide-ranging significance. The researchers now plan to explore how DHX29 affects health and disease.
“We have long been fascinated by how cells interpret the hidden layer of information embedded within the genetic code, so discovering the molecular factor that allows human cells to read and respond to this hidden code has been particularly rewarding,” says team leader Osamu Takeuchi.
Reference: “Human DHX29 detects nonoptimal codon usage to regulate mRNA stability” by Fabian Hia, Yitong Wu, Masanori Yoshinaga, Sakurako Goto-Ito, Wakana Iwasaki, Koshi Imami, Hirotaka Toh, Peixun Han, Ting Cai, Takayuki Ohira, Akira Fukao, Daron M Standley, Yuichi Shichino, Masaki Takegawa, Toshinobu Fujiwara, Tsutomu Suzuki, Shintaro Iwasaki, Michael C. Bassik, Takuhiro Ito and Osamu Takeuchi, 19 March 2026, Science.
DOI: 10.1126/science.adw0288
Never miss a breakthrough: Join the SciTechDaily newsletter.
Follow us on Google and Google News.
